We wrapped up the last day with age-depth models and independent project presentations.

3 classical age models include linear interpolation (connect the dots), regression (linear or polynomial), and smooth spline. If you use classical age models, CLAM (classical age models) is an R package that is good step forward.
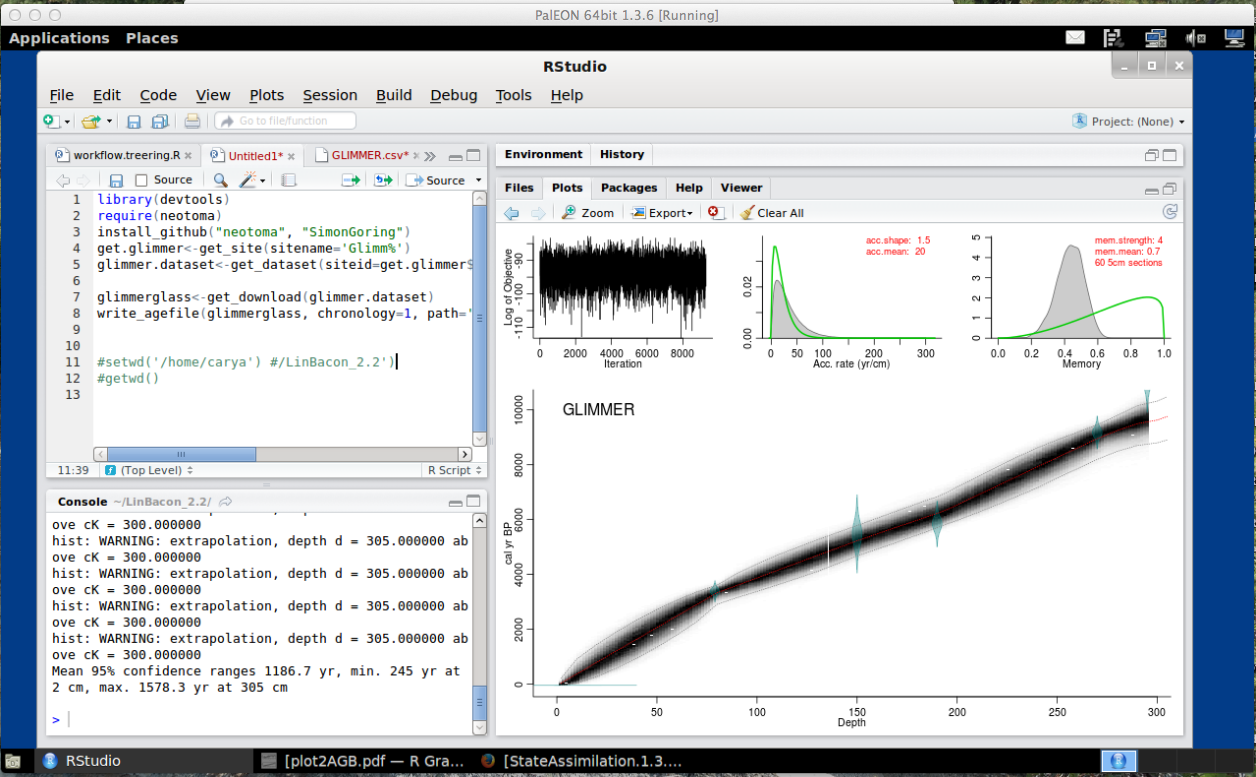
The field is moving away from classical models and starting to use Bayesian Age Models (started in the mid-1990s). The age model we discussed and applied to actual data is BACON.
Here is an example of the age model example the class walked through for sediment collected from Glimmerglass Lake. It is a nice example of a model that is well mixed, fits well, and goes through all radiocarbon dates.
The afternoon was dedicated to student presentations on their independent projects. There were a wide range of independent project topics including:
- State space model for plant macrofossils
- Quantify variance between lake pollen loadings using observed pollen proportions
- 2000 years of inferred forest structure in the southern Sierra Nevada from a wet meadow
- Empirical succession model application to the fading record problem and uncertainty in forest density trajectories
- Using pollen and squirrel fossils to explore two datasets with irregular spacing of time
- Exploring Colombian vegetation using 4 proxies: 13C, volcanic minerals, archaeological artifacts and charcoal
- Fire signals in southern CA
- Pollen in Australian lakes
- MCMC of Alaska Picea glauca (white spruce)
- Detecting dust storms in lake sediment cores – are the dust storms increasing?
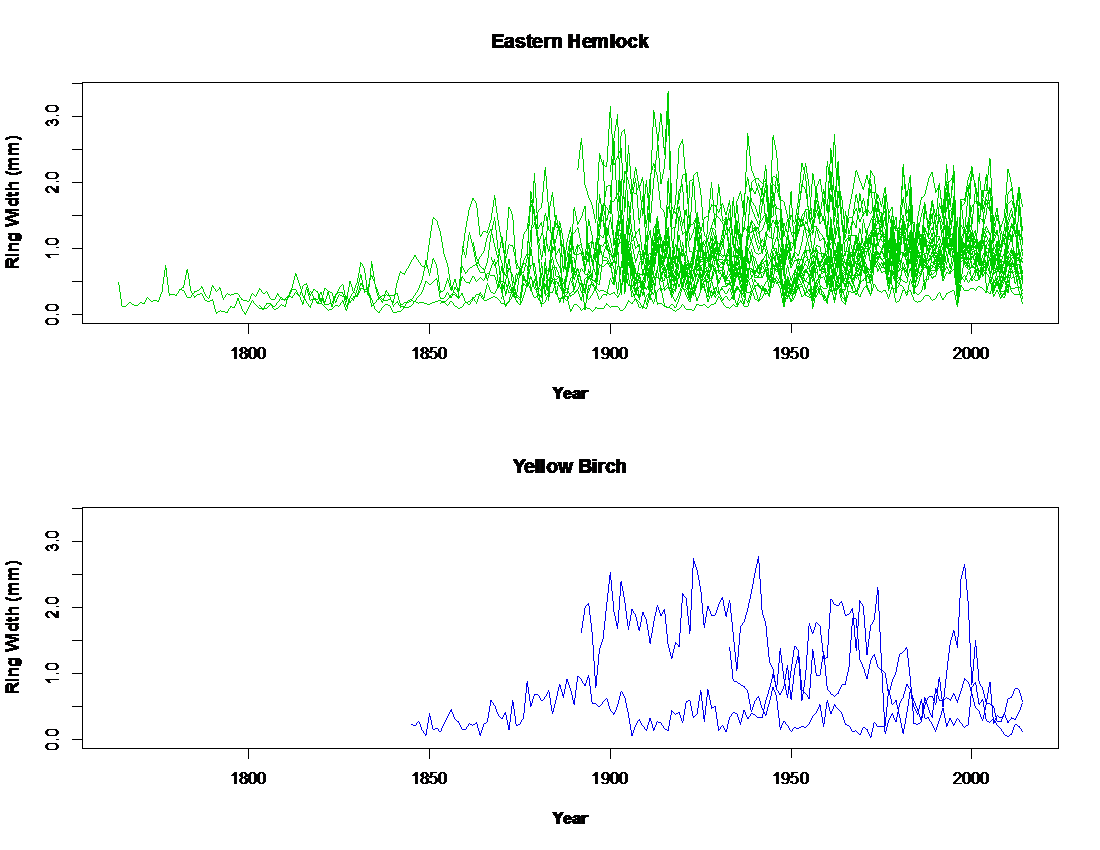
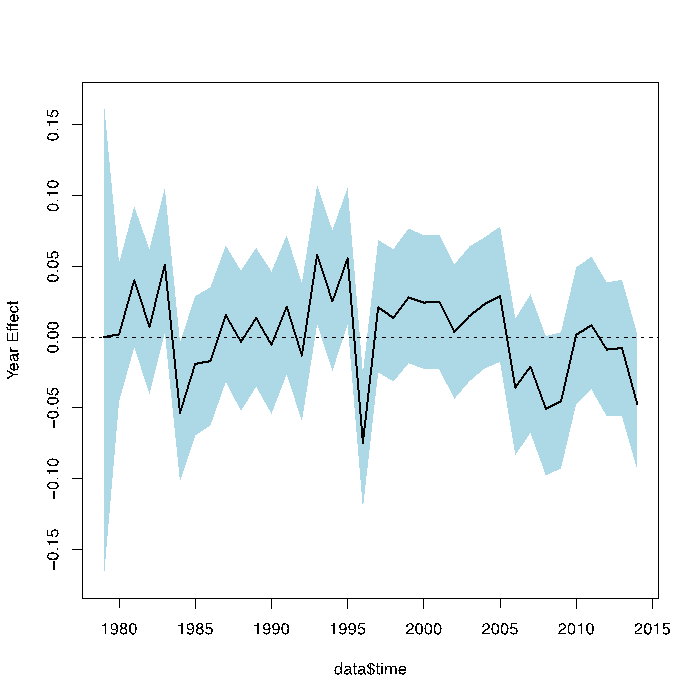
- Using eddy flux data and terrestrial models to estimate NPP and biomass from tree rings
For only having 6 hours work, the projects turned out really well, with lots of converged iterations. It was great to see how much the students were able to pick up over the past 6 days.
We wrapped up big week with lots of hard work by celebrating with a classic Friday night fish fry dinner at Bent’s Camp!