News
What are the “design rules” for engineering multicellular tissues?
We use a combination of experimental and computational approaches to reverse-engineer pattern formation and tissue morphogenesis, growth control, and regeneration to inspire new ways to treat human diseases, such as cancer, and to accelerate wound healing and regeneration. Our research leads to the discovery new targeting approaches for treating a broad range of diseases and a better understanding of the rules governing organ development.
Lab openings:
We are looking for talented and motivated graduate students and post-doctoral researchers with interests in two areas:
1. Synthetic, quantitative biology of multicellular systems; and
2. Computational modeling of cell signaling or systems identification.
Interested undergraduates and potential graduates and post-doctoral researchers are welcome to inquire about available positions in the lab. Please note that undergraduate research positions require a 12 hour per week commitment to the research project.
2025
June 2025
(06/20): Congrats Benjamin for another milestone accomplished! He just passed his candidacy proposal exam. So proud of you! Celebrated this legend at Legends of Notre Dame.

(06/05): George, welcome to MSEL!
(06/04): Welcome Stacey to MSEL!
May 2025
(05/18): Congrats Dr. Mim! Graduating her PhD and now off to her post-doc position at Northwestern University Feinberg School of Medicine. We are so proud of you!

April 2025
(04/27): Welcome Naomi to our group!
Celebrating birthdays and Naomi’s welcoming to MSEL in Bonefish Grill! We had a blast catching up and celebrating everyone.


(04/14): Congratulations to everyone who participated in the Thirteenth Annual Harper Cancer Research Institute Cancer Research Day!





(04/11): Congratulations, David on winning the BIPH 2025 O’Brien Endowment for Excellence Graduate Fellowship!
(04/11): The Q-Bio@ND-EMBRIO Joint Symposium was a success, with many networking opportunities, connecting across different programs and learning great quantitative work from our peers.
Click this link to go to their webpage!

(04/02): Congratulations to David on his successful Candidacy exam! One more step towards your PhD!
2024
December 2024
Celebrating the holiday season is always a fun time at MSEL!

November 2024
Congratulations, Dr. Mayesha S. Mim! We are so proud of you for defending your dissertation. We are so glad we got to celebrate Mayesha while enjoying each other’s company, and of course, hot pot!

Welcome Caitlin to our group!
September 2024
Welcome Yikang to our group!
August 2024
Welcome Beatriz to our group!
May 2024
We are excited to announce that our new collaborative research paper, highlighted here, is now available online! This study significantly advances our understanding of the biomechanical interactions defining the rules governing organ growth by generating and testing a “digital twin” of the maturing fruit fly wing disc. We thank our collaborators Prof. Mark Alber and Prof. Weitao Chen, and special acknowledgment is noted to the authors who performed the research: Dr. Nilay Kumar, Jennifer Rangel Ambriz, Dr. Kevin Tsai, Mayesha Mim, and Dr. Marycruz Flores-Flores.
Funding support statement: This work was supported by the National Science Foundation grant NSF2029814 to all co-authors and by a pilot grant from the Northwestern University NSF-Simons Center for Quantitative Biology to JZ. The Zartman group was supported by EMBRIO Institute, contract #2120200, a National Science Foundation (NSF) Biology Integration Institute, and NIH Grant R35GM124935 (JZ).
2023
July 2023
We are organizing the 2nd Annual EMBRIO training workshop at ND.
May 2023
It was wonderful to connect with Dr. Vijay Velagala at graduation
April 2023
Dr. Nilay Kumar successfully defended his dissertation. Excellent job!
March 2023
We welcome Dr. Pablo Cisternas and Dr. Marycruz Flores Flores to our lab.
February 2023
Dr. Vijay Velagala defended his dissertation. Way to go!
2022
October 2022
We are organizing one-day virtual workshop on “Reverse Engineering Mechanisms of Morphogenesis” with invited speakers from Notre Dame and Mexico to seed new multidisciplinary collaborations and exchange between our departments and institutions.

JUNE 2022
Research in the summer is the best! Especially with an occasional ice cream break.

We welcome new students into the group:
Lauren Hawthorne is joining our team as she embarks on her Bioengineering Ph.D. studies.
Anne Malott is working with Mayesha on an exciting project related to cancer through the ND HCRI’s Research Cures Cancer Corps “Scholar.”
Jose Martinez Alvarado is joining us for a summer research experience through SROP 2022. He is from the University of Puerto Rico, Mayaguez, Department of Industrial Biotechnology.
Marycruz Flores Flores , a Ph.D. student in the lab of Marcos Nahmad at Cinvestav is collaborating with us too.
Welcome Lauren, Anne, Mary, and Jose!
May 2022
The Multicellular Systems Engineering Research Lab (https://t.co/PlHbfnAcWe) has once again proven that they can get their PI out of a tight spot & in under an hour! Way to go, Team! @NDCBE @NDBioeng pic.twitter.com/g8EuPsTbh1
— Jeremiah Zartman (@JeremiahZartman) May 13, 2022


If we can escape and defuse a dangerous situation with 55 seconds to spare, we are ready to tackle any research question as a team!

Food and Fun! A great way to wrap up the semester and gear up for a great summer of research
Well done on Vijay on your talk at the Annual Midwest Imaging symposium!
Congratulations to Francisco for completing his candidacy exam. A very important milestone!
April 2022
We are recruiting students for Ph.D. projects!
 Happy Birthday, Mayesha! Research is always fun with a little cake and birthday celebration added.
Happy Birthday, Mayesha! Research is always fun with a little cake and birthday celebration added.
Congratulations to Giorgia Giordano on successfully defending her research thesis! We are proud fo Giorgia and wish her the very best as she graduates this spring!
March 2022
Congratulations to Dr. Dharsan Soundarrajan on successfully defending his dissertation! We are proud of him and we wish him the very best in his future endeavors! pic.twitter.com/BZRvv2lYd3
— Jeremiah Zartman (@JeremiahZartman) March 29, 2022

2021
DECEMBER 2021
Sometimes it is good to relax and enjoy good company. A recent lab get-together.

November 2021
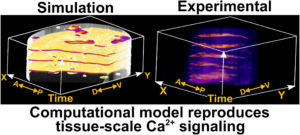
Congratulations to Dharsan and Francisco on their published study in PLOS Computational Biology!
October 2021
Congratulations to Vijay on his publication in Biophysical Journal!
SePTEMBER 2021
We are a part of the new Biology Integration Institute (Emergent Mechanisms in Biology of Robustness Integration and Organization Institute (EMBRIO) !
We welcome David Gazzo to our lab. Welcome, David!
Enjoying a nice autumn evening and lab celebration!

July 2021
Congratulations to Dr. Megan Levis for defending her dissertation! Well done, Megan!
We also welcome Marycruz Flores, as an international visitor and collaborator to our research group for an exciting research collaboration that is partly supported by ND International. She is a researcher advised by Dr. Marcos Nahmad at Cinvestav.
2019
February 2019
Congratulations to Dharsan Soundarrajan for passing his Ph.D. candidacy exam!
Collaborative work with Saverio Gentile entitled “Potassium channel activity controls breast cancer metastasis by affecting β-catenin signaling” is now published in Cell Death and Disease!
January 2019
Our work on decoding calcium signaling dynamics at the organ-level is accepted published in Biophysical Journal! The work is entitled, “Decoding Calcium Signaling Dynamics during Drosophila Wing Disc Development.”Way to go everyone!
2018
October 2018
We officially welcome Francisco Huizar, who is pursuing a Bioengineering Ph.D. program, to the lab!
August 2018
Congratulations to Dr. Pavel Brodskiy on his successful dissertation defense. Well done!
July 2018
Our short perspective on the new field of synthetic developmental biology is highlighted at Science Trends.
Ulises, Ana, Mark and Anthony all provide research updates at group meeting. It was great having you guys join our group for the summer!
Summer lab picnic: Lots of fun!

May 2018
Dr. Zartman is promoted to Associate Professor (effective July 2018).
January 2018
Dr. Zartman named as a 2018 Rising Star by the BMES Cellular and Molecular Bioengineering Special Interest Group working group.
2017
December 2017
We welcome Mascha and Ramezan to our group! Welcome
October 2017
We welcome Nilay and Vijay to our group! Welcome!
October 2017
October 6, 2017: Congratulations to Cody Narciso for successfully defending his dissertation. Way to go, Dr. Narciso!
September 2017
Current: We are looking for talented post-doctoral researchers with experience in either two areas:
1. Molecular cell biology and advanced Drosophila genetics
2. Computational modeling of cell signaling or systems identification
We are also looking for lab technicians with expertise in Drosophila and cell biology. Please contact Professor Zartman directly.
August 2017
We have been awarded a NIH R35 grant entitled: “Regulation and Function of Multicellular Calcium Signaling in Epithelial Growth and Regeneration.” In this research program we seek to discover the underlying principles of integrative cell communication mediated by calcium signaling.
A long-term goal of this research is to learn how to manipulate calcium signaling to control cellular functions such as growth, motility, or death. Such knowledge will inspire methods to regulate cell differentiation for stem cell engineering applications or to stop metastatic cancers.
July 2017
Our work “Release of Applied Mechanical Loading Stimulates Intercellular Calcium Waves in Drosophila Wing Disc” in Biophysical Journal is now online. This work was highlighted as a “New and Notable” by the journal!
July 17, 2017: Our work with the Alber, Goodson and Weisel groups to identify efficient tissue clearing method for whole blood clots is now published at Biomedical Optics Express! This work has been highlighted at EurekAlert and Science Daily.
July 10, 2017: Our collaborator Jochen Kursawe (Oxford University) visited us and also gave a seminar talk: Quantitative approaches to investigating epithelial morphogenesis
April 2017
April 8, 2017: We are co-organizing the 5th Midwest Q-Bio Symposium @ University of Notre Dame. It was a great turnout!
March 2017
March 2017: Contratulations to Paulina Eberts for her National Science Foundation Gradute Fellowship award!
Paulina was selected as an undergraduate student in chemical engineering. Her thesis is “Extension of Non-destructive 3D Immunostaining to Paraffin Embedded Tissues.”
2016
November 2016
November 2016: Our collaborative work is now published in Interface! Robust cell tracking in epithelial tissues through identification of maximum common subgraphs. Congratulations especially to lead author Jochen Kursawe.
November 2016: We welcome two new members to the lab: Jamison Jangula and Dharsan Soundarrajan.
July 2016
July 2016: Congratulations to Pavel! He was selected as an Walther Cancer Foundation Interdisciplinary Interface Training Project (IITP) fellow through the Harper Cancer Research Institute.
June 2016
June 7, 2016: Congratulations, Dr. Miranda Burnette! Great job on your dissertation defense. Way to go!
May 2016
May 2016: Megan, welcome to the lab! Megan Levis joins the group.
March 2016
March 1 2016: We have been awarded an NSF CAREER award for our project, “Integrative Analysis for Reverse Engineering Embryonic Pattern Repair Mechanisms.”
March 1 2016: In collaboration with the Hoelzle and Zhang labs, a short communication describing on-chip 3D tissue histology for microbiopsies is now online in Biomicrofluidics. Great work team! Links to the press release is found here.
February 2016
February 26, 2016: Congratulations to Qinfeng Wu for passing his Ph.D. Candidacy exam. Way to go!
2015
December 2016
A collaborative study with the Fletcher and Baker groups on the capabilities and limitations of tissue size control through passive mechanical forces is now in press in PLOS Computational Biology!
August 2015
Our study examining the spatiotemporal dynamics of calcium flashes due to local wounding is now online.
July 2015
July 2015: Congratulations to Cody Narciso, who was named a 2015/16 Berry Family Foundation Graduate Fellowships in Advanced Diagnostics & Therapeutics. See more at here.
April 2015
April 2015: Congratulations to Qinfeng Wu on winning the HCRI Research Like a Champion Award!
From the CBE Website: “Qinfeng Wu, a graduate student in the Department of Chemical and Biomolecular Engineering, was one of the 2015 winners in the annual Research Like a Champion competition. Sponsored by the Harper Cancer Research Institute, the competition is open to individual students/student groups across all disciplines of the University. Participants submit innovative approaches to the fight against cancer. The submitted proposals address causes, treatment, or prevention of the disease.
Wu’s winning project, “Cell Competition in Breast Cancer: Functional Identification of Genes that Determine Whether Breast Cancer Cells Out-compete Breast Epithelial Cells,” focuses on how breast cancer cells avoid tumor suppression treatments and beat out normal cells for nutrients, as well as how best to identify and change the mechanisms through which this cell competition occurs.”
April 2015: Congratulations to the start-up Enlightened Diagnostics, Inc. on their achievements at the McCloskey Business Plan competition and at the Rice Business Plan Competition. For a description of their success, click here.
February
February 2015: Congratulations to the Englightened Diagnostics team on their victory at the Cardinal Challenge business plan competition! The team is presenting on a novel 3-D imaging platform developed iin collaboration with the Hoelzle and Zhang labs to improve the amount of information available for clinicians from biopsies.
January
January 2015: Cody passed his Candidacy exam. Congratulations!
2014
July
July 2014: Miranda publishes her chemically defined medium for long-term maintenance and growth of Drosophila cells. The first truly chemically defined medium for Drosophila cell culture. In addition, the paper highlights a strategy for rationale media design. Congratulations!
2014
April 2014
April 15, 2014: Cody Narciso and Kyle Cowdrick (Mentor: Siyuan Zhang) win the Harper Cancer Research Institute’s“Research Like A Champion” Award. Congratulations!
January
January 30, 2014: Miranda passes her Candidacy exam. Congratulations!
2013
December
December 12, 2013: Lab Christmas party. Merry Christmas and Happy New Year!
November
Pavel Brodskiy and Qinfeng (Austin) Wu officially join the group. Welcome!
June
June 11, 2013: Jessica Freeman selected as a HCRI Summer Undergraduate Research Fellow. Congratulations Jessica!
March
March 4, 2013: Cody Narciso presented at the Workshop on Physical Approaches to Studying the Cytoskeleton and Cell Motility: https://www3.nd.edu/~icsb/Workshop_2013/.
January
January 12-16, 2013: Miranda presented a poster at SLAS 2013: Miranda Burnette, Jonathan Chen, and Jeremiah Zartman, Combinatorial Inverse Drug Screen Pipeline to Probe Epithelial Wound Healing in a Model Genetic Organ Culture System. SLAS 2013. January 12-16, 2013.
2012
October
October 2012: Cody Narciso joins the lab. Welcome!
March
March 2012: Teresa Brito-Robinson joins the lab. Welcome!
January
January 2012: Lab officially opens. Miranda Burnette joins the lab. Welcome!
Events
Upcoming events:
To be updated.
Past events:
Simplified microfluidics and bioprinting systems for research and education.
Fernando Ontiveros,
PhD. Assistant Professor Biology Department, St. John Fisher College
Tuesday, August 8, 4-5 pm
McCourtney 241
Widespread adoption of novel research technologies is often hampered by cost and lack of equipment and expertise. Overcoming these hurdles can prove challenging. For example, improved protocols that produce reliable results may manage to reduce production time and cost of materials, but may still require procedures and/or the use of equipment not commonly available to most research and teaching laboratories. On the other hand, the quality and/or reproducibility of results obtained using low cost approaches may be compromised by the use of unreliable materials. Our goal is to provide research groups and educational settings with microfluidics and bioprinting systems that are both reliable and affordable. In the area of microfludics we describe a type of device that makes exclusive use of consumer-grade components and equipment. The user-designed devices consist of as little as three layers of a polymer film, with microchannels shaped by an inexpensive craft cutter, and sealed by thermal lamination. The chips are flexible and ten to thirty-times thinner than the common PDMS/ glass alternative. Fabrication time is in the order of minutes (allowing for rapid iteration), and the method requires very little training. In the area of bioprinting we make use of a consumer-grade 3D printer fitted with a heated nozzle which delivers a biocompatible hydrogel that sets as it comes into contact with a cooled surface. Our objective is to produce scaffolds in a variety of shapes that promote 3-dimensional arrangements of cells in culture. In summary, our systems lower the barrier-to-entry for robust microfluidic devices and biocompatible tissue scaffolds, facilitating the adoption of formerly out-of-reach technologies.
Guest speaker
Monday, July 10 3 – 4 PM,
McCourtney Hall Conference Room 203
Quantitative approaches to investigating epithelial morphogenesis
Jochen Kursawe, University of Oxford, Oxford
Recently, the experimental study of morphogenesis has thrived due to a rise in quantitative methods. The resulting avalanche of quantitative data requires us to rethink the scientific method. We need to design quantitative hypotheses through mathematical models, make quantitative experimental predictions, devise methods for quantitative data analysis, and design methods for quantitative inference using models and data. This work aims to enable this transition for the integrative analysis of morphogenesis in epithelia. We conduct the first systematic numerical analysis of a widely used cell-based model of epithelia, the vertex model, and estimate to what extent quantitative model predictions may be influenced by parameter values and implementation details. We then apply this model to a key question in developmental biology by constructing a quantitative theory for tissue size control in the embryonic epidermis of the fruit fly Drosophila, using the model to predict the outcomes of future experiments. We further devise a method for estimating mechanical parameters of vertex models from imaging data and quantifying the uncertainty associated with such estimates. Finally, we propose a novel algorithm for robust cell tracking in live-imaging microscopy videos of epithelial tissues that illustrates how graph theoretic concepts may be used to overcome challenges in quantitative data analysis. Together, these contributions will enable the quantitative study of epithelia for a wide range of applications.
Christmas dinner with the lab group – December 6, 2012 at J.W. Chen’s. Lot’s of fun!
Lab Picnic was held on May 2013.